The Problem
- Standard allergy tests (IgE, skin prick) often yield ambiguous results and can’t predict true allergic reactions reliably.
- The gold-standard oral food challenge is risky, resource-intensive, and used sparingly, leaving many patients undiagnosed or in limbo.
- Rising food allergy rates heighten the need for better diagnostics to distinguish truly allergic patients without putting them through dangerous procedures.
The Solution
- Built an ML model that combines multiple inputs (specific IgE levels, skin test results, patient history) to predict if a patient is truly allergic or likely tolerant.
- Trained on real hospital data (patients with known oral challenge outcomes) so the model “learns” the patterns of feature combinations that signal a true allergy.
- Designed a prototype pipeline (“ML-Allergy”) integrated with the EHR: it pulls a patient’s lab results and history, runs the ML risk algorithm, and outputs a risk score with guidance to clinicians.
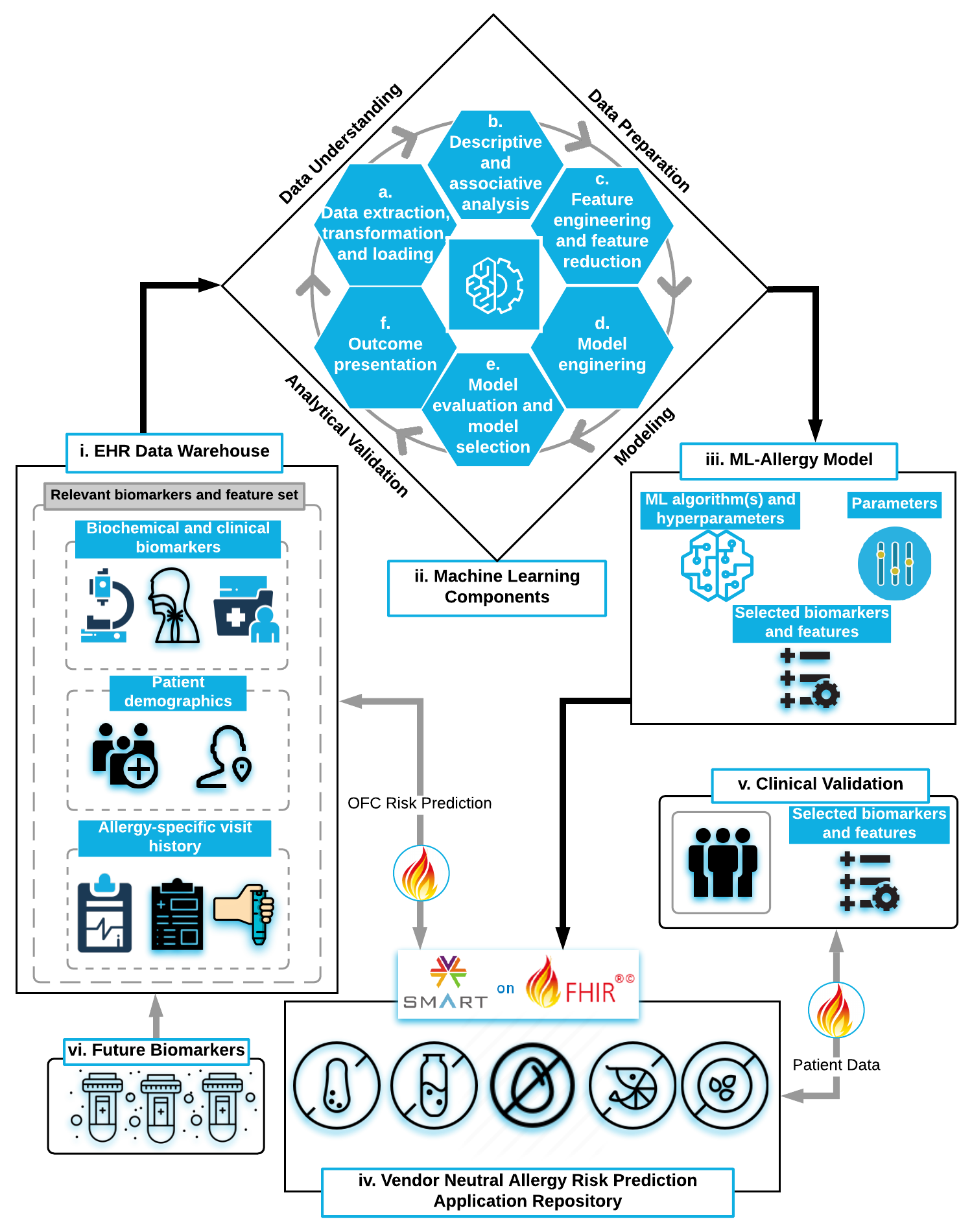
Architecture Overview
- Six-component ML pipeline from data extraction to decision support. It automatically pulls relevant lab results and clinical history from the EHR data warehouse via ETL processes.
- Feature engineering handles ~20+ predictors (IgE, ratios, symptoms, etc.), accommodates missing data, and uses regularization to rank important features and avoid overfit.
- Iterative model training with cross-validation tests various algorithms (logistic regression, random forest, gradient boosting) to maximize predictive AUC while maintaining generalizability.
- The final model achieved high performance (AUC ~0.96) and was configured to emphasize safety (e.g., tuning thresholds to minimize false negatives for allergies).
- The output is presented in a clinician-friendly format within workflow: a risk score (“High risk – 85% chance of reaction”) along with key contributing factors, seamlessly delivered alongside existing lab reports.
Results and Impacts
- The ML risk model reached ~96% accuracy (AUC ~0.96) in validation, far exceeding the predictive power of any single test, which translates to much more confident clinical decision-making.
- Combining multiple test results proved its value: the model clearly separated allergic vs. tolerant patients, confirming that multiple weak indicators together create a strong signal.
- In practice, this enables doctors to avoid unnecessary and risky food challenges for low-risk patients and to focus resources on those truly at risk, improving safety and reducing patient anxiety.
Skills and Tools Used
| Technique/Skill | Tools/Implementation |
|---|---|
| Skill/Tool Category | Application in ML-Allergy — Food Allergy Risk Stratification |
| Data Collection (EHR) | SQL and Python (Pandas) to extract and merge allergy test results and clinical notes from hospital databases (Epic EHR) |
| Machine Learning (Python) | Scikit-learn & LightGBM for model development; cross-validation, grid search, and regularization (L1/L2) for feature selection and tuning |
| Statistical Analysis | ROC/AUC analysis, bootstrapped confidence intervals, and custom threshold tuning to maximize negative predictive value (safety first) |
| Clinical Domain Integration | Incorporated medical expertise (e.g., weighting “history of anaphylaxis” appropriately); close collaboration with allergists to embed domain logic in the model |
| Healthcare Data Standards | Used ICD-10 and LOINC codes to identify data in EHR; leveraged FHIR resources to integrate the model output into clinical systems (ensuring compatibility and privacy) |
| Communication & Visualization | Presented results in clinician-friendly terms (e.g., “avoid X% of unnecessary challenges”) and used clear visual aids (calibration plots, decision curves) to gain physician buy-in |
Cross-Project Capabilities
- Clinical Decision Support Development: End-to-end experience building a clinical ML tool (data ingestion, model, workflow integration) carried into later projects like ICU decision support and maternal-infant care tools.
- Interdisciplinary Collaboration: Skill in partnering with clinicians (allergists in this case) to ensure ML solutions meet real-world needs – later applied with ophthalmologists (Vision project) and intensivists (ICU project).
- Regulatory/Ethical ML Practice: Navigated patient data privacy (HIPAA, IRB approvals) and clearly communicated model limitations to stakeholders – a critical skill in all healthcare AI projects dealing with sensitive data.
Published Papers/Tools
- Internal White Paper: “ML Composite Biomarker Panel for Food Allergy Diagnosis” – detailed proposal circulated within the hospital to outline the project’s design and rationale.
- Hospital Knowledge Sharing: Findings presented at Boston Children’s Hospital grand rounds and an innovation showcase, raising awareness of ML’s potential in diagnostics.
- Prototype Tool: Developed a pilot “Allergy ML Risk Calculator” (Excel interface + Python backend) for clinicians to input patient data and get a risk estimate, now undergoing evaluation for integration into the EHR’s decision support system.