The Problem
- Dangerous drops in blood oxygen (hypoxemia) can occur between intermittent arterial blood gas tests, since these invasive checks are done only a few times a day.
- Pulse oximeter readings (SpO₂) are continuous but become unreliable at the top end (97–100% SpO₂) – they can’t distinguish a safe patient from one with dangerously high oxygen levels.
- Existing surrogate formulas for PaO₂ are often inaccurate, especially at saturation extremes, and don’t adapt to individual patient differences, making clinicians hesitant to trust them.
The Solution
- Collected a large dataset of paired ABG measurements and bedside monitor readings (~52,000 samples) from ICU patients, and used it to develop better estimation models.
- Explored machine learning approaches (including neural networks using inputs like recent SpO₂ trends and heart rate) and also optimized a conventional oxygenation equation by tuning physiological parameters to fit real patient data.
- Selected a new empirically tuned equation (“Sauthier ePaO₂”) that provided the best accuracy. Implemented it for continuous use: the ICU monitor (or a linked computer) takes in streaming SpO₂ and heart rate and computes an estimated PaO₂ every minute, displaying it to clinicians and triggering alerts if oxygen drops.
- Built in real-time quality control: the algorithm watches the difference between pulse oximeter heart rate and EKG heart rate as a noise indicator – if they diverge, it treats the SpO₂ data as unreliable and adjusts or flags accordingly.
Architecture Overview
- Data Pipeline: Retrieved high-frequency vital signs data around each ABG draw from two hospitals’ systems (merging data logged every 5 seconds from one system and every 1 second from another, standardized to 5-second intervals). Stored all combined data in a PostgreSQL database for analysis.
- Analysis & Modeling Environment: Used Python and R extensively (Pandas, NumPy, SciPy, scikit-learn, Keras/TensorFlow) to clean data, engineer features (e.g., recent SpO₂ averages, slopes, heart-rate disparities), and train/test multiple candidate models.
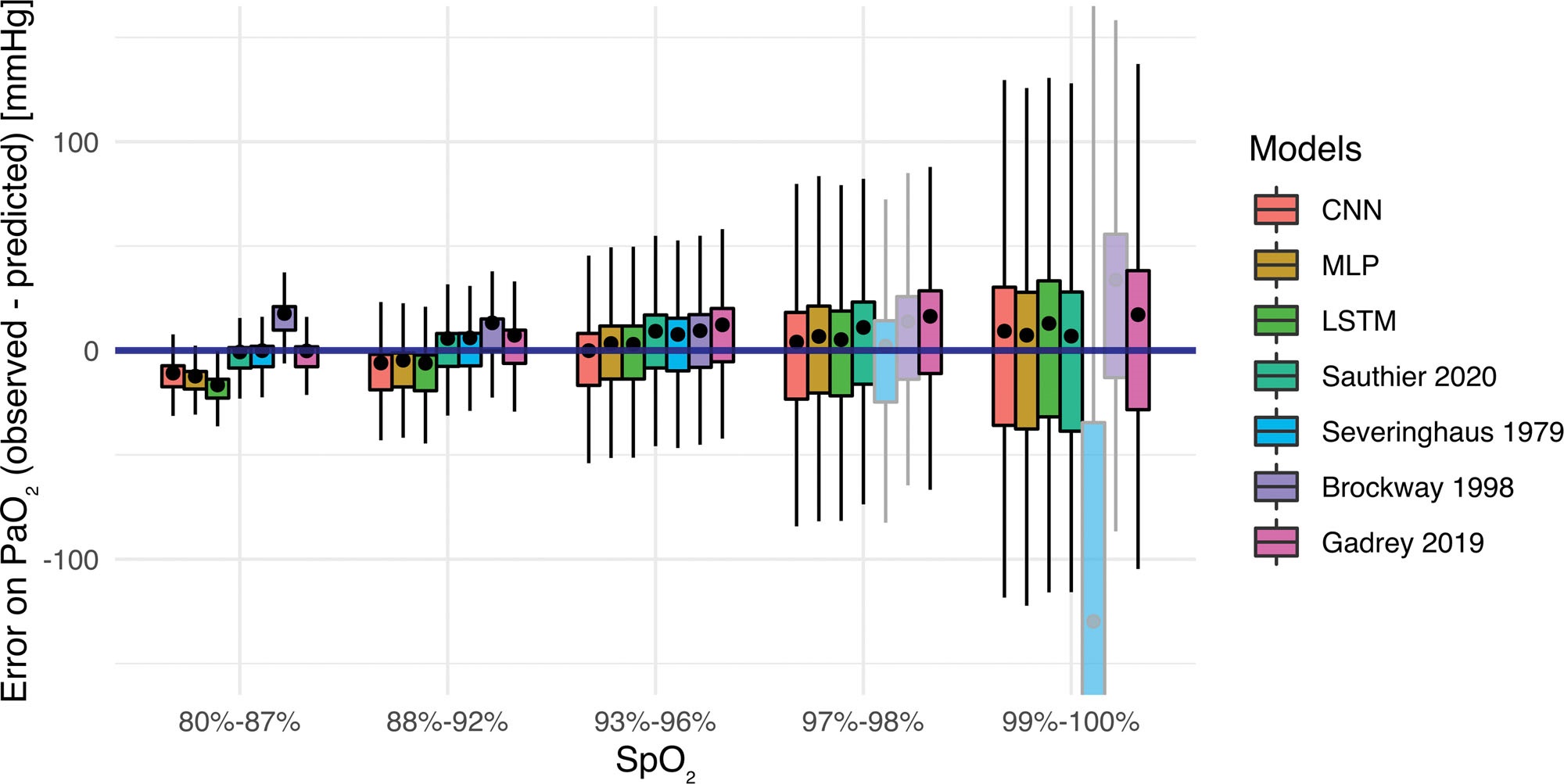
- Model Evaluation: Employed rigorous statistical evaluation – generated Bland-Altman plots to assess agreement between estimated and actual PaO₂, calculated intraclass correlation coefficients (agreement), and examined error across different SpO₂ ranges to ensure reliability even at high saturations.
- Deployment Design: Specified how the final algorithm would run on existing ICU monitors: a lightweight script or plugin continuously ingests real-time vitals, applies the estimation formula, and outputs a refreshed PaO₂ estimate every few seconds. This can feed into the bedside monitor display (trend graphs, ARDS index calculations) and alarm systems for immediate clinical use.
- Safety Considerations: Built into the design fail-safes like capping displayed values beyond validated ranges (e.g., instead of an implausible number at extreme highs, show an indicator like “PaO₂ very high”) and ignoring data during sensor disconnections or extreme noise, ensuring the tool would be trusted by clinicians.
Results and Impacts
- The optimized estimator provided sensible, stable PaO₂ readings even when SpO₂ was 100%, overcoming the “infinite reading” problem of traditional equations and giving clinicians meaningful data at the high end of oxygenation.
- Continuous monitoring enabled earlier detection of hypoxemia trends – for example, the system could alert if a patient’s oxygen was quietly deteriorating between blood gas tests, allowing interventions (like ventilator adjustments) sooner and improving ARDS management by tracking severity in real time.
- The work was published in Critical Care Explorations (2021) and has since been cited in multiple studies, influencing the field of digital critical care. It provided a blueprint for integrating predictive algorithms into ICU workflows, with discussions underway to pilot the estimator in a real ICU setting.
Skills and Tools Used
| Technique/Skill | Tools/Implementation |
|---|---|
| Skill/Tool Category | Application in ICU Clinical Decision Support – Continuous Hypoxemia Monitoring |
| Data Engineering (ICU) | SQL (PostgreSQL) and Python to manage ~65k blood gas records with high-frequency vital sign data; automated data extraction around events |
| Statistical Analysis | Advanced stats: Bland-Altman agreement analysis, bootstrapped confidence intervals, hypothesis tests to validate model accuracy and bias |
| Machine Learning & Modeling | Keras/TensorFlow to prototype neural networks for regression; exhaustive parameter search to optimize a physiological equation for best fit |
| Domain Knowledge (Critical Care) | Integrated understanding of oxyhemoglobin dissociation curves, ARDS criteria, and physiological factors (P50 shifts) to guide modeling and interpret results |
| Software Development | Python scripting for simulating real-time monitoring (feeding sequential data to the model); implemented the final formula in an efficient form suitable for bedside device integration |
| Collaboration & Publication | Coordinated a multi-center team (Boston & Montreal) under data-sharing agreements; co-authored a peer-reviewed journal paper and produced publication-quality figures (e.g., error distribution charts) |
Cross-Project Capabilities
- Translating data science research into a clinical tool: experience in going from retrospective analysis to a validated model ready for real-world use (as done here) is directly applicable to other projects where ML models must be deployment-ready (e.g., the allergy risk model or mother-infant dashboard logic).
- Time-series data expertise: skills in handling and analyzing high-frequency time-series (ICU monitor streams) are transferable to other domains, including real-time social media feeds in outbreak surveillance or sensor data in IoT health projects.
- Multi-site data integration: learned to standardize and combine data across different hospital systems, a capability that proved valuable in later projects focused on data interoperability (such as linking records in the mother-infant health information exchange project).
Published Papers/Tools
- Journal Publication: Critical Care Explorations (Oct 2021) – “Estimated PaO₂: A Continuous and Noninvasive Method to Estimate PaO₂ and Oxygenation Index.” This open-access paper details the methodology and validation results, serving as the primary reference for this work. Paper
- Open Science Resources: The team shared portions of the de-identified dataset and analysis code via an online repository referenced in the publication, enabling other researchers to reproduce or build upon the findings.
- Community Impact: The method has been discussed in critical care forums and was highlighted in a critical care innovation newsletter, indicating its relevance. Its successful publication and citations (including by a 2024 digital medicine article) underscore its influence on subsequent research in noninvasive monitoring.